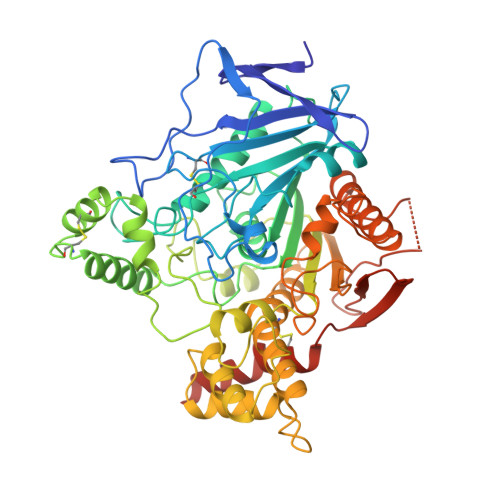
Crystal structures of aged phosphonylated acetylcholinesterase: nerve agent reaction products at the atomic level.
Millard, C.B., Kryger, G., Ordentlich, A., Greenblatt, H.M., Harel, M., Raves, M.L., Segall, Y., Barak, D., Shafferman, A., Silman, I., Sussman, J.L.(1999) Biochemistry 38: 7032-7039
- PubMed: 10353814
- DOI: https://doi.org/10.1021/bi982678l
- Primary Citation of Related Structures:
1CFJ, 1SOM, 2DFP - PubMed Abstract:
Organophosphorus acid anhydride (OP) nerve agents are potent inhibitors which rapidly phosphonylate acetylcholinesterase (AChE) and then may undergo an internal dealkylation reaction (called "aging") to produce an OP-enzyme conjugate that cannot be reactivated. To understand the basis for irreversible inhibition, we solved the structures of aged conjugates obtained by reaction of Torpedo californica AChE (TcAChE) with diisopropylphosphorofluoridate (DFP), O-isopropylmethylphosponofluoridate (sarin), or O-pinacolylmethylphosphonofluoridate (soman) by X-ray crystallography to 2.3, 2.6, or 2.2 A resolution, respectively. The highest positive difference density peak corresponded to the OP phosphorus and was located within covalent bonding distance of the active-site serine (S200) in each structure. The OP-oxygen atoms were within hydrogen-bonding distance of four potential donors from catalytic subsites of the enzyme, suggesting that electrostatic forces significantly stabilize the aged enzyme. The active sites of aged sarin- and soman-TcAChE were essentially identical and provided structural models for the negatively charged, tetrahedral intermediate that occurs during deacylation with the natural substrate, acetylcholine. Phosphorylation with DFP caused an unexpected movement in the main chain of a loop that includes residues F288 and F290 of the TcAChE acyl pocket. This is the first major conformational change reported in the active site of any AChE-ligand complex, and it offers a structural explanation for the substrate selectivity of AChE.
Organizational Affiliation:
Department of Neurobiology, Weizmann Institute of Science, Rehovot, Israel. [email protected]