The discovery of novel benzofuran-2-carboxylic acids as potent Pim-1 inhibitors.
Xiang, Y., Hirth, B., Asmussen, G., Biemann, H.P., Bishop, K.A., Good, A., Fitzgerald, M., Gladysheva, T., Jain, A., Jancsics, K., Liu, J., Metz, M., Papoulis, A., Skerlj, R., Stepp, J.D., Wei, R.R.(2011) Bioorg Med Chem Lett 21: 3050-3056
- PubMed: 21507633
- DOI: https://doi.org/10.1016/j.bmcl.2011.03.030
- Primary Citation of Related Structures:
3R00, 3R01, 3R02, 3R04 - PubMed Abstract:
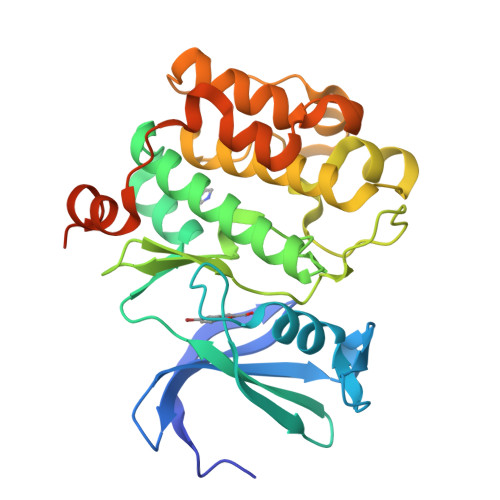
Novel benzofuran-2-carboxylic acids, exemplified by 29, 38 and 39, have been discovered as potent Pim-1 inhibitors using fragment based screening followed by X-ray structure guided medicinal chemistry optimization. The compounds demonstrate potent inhibition against Pim-1 and Pim-2 in enzyme assays. Compound 29 has been tested in the Ambit 442 kinase panel and demonstrates good selectivity for the Pim kinase family. X-ray structures of the inhibitor/Pim-1 binding complex reveal important salt-bridge and hydrogen bond interactions mediated by the compound's carboxylic acid and amino groups.
Organizational Affiliation:
Department of Medicinal Chemistry, Drug and Biomaterial R&D, Genzyme Corp., 153 Second Avenue, Waltham, MA 02451, USA. yibin.xiang@genzyme.com