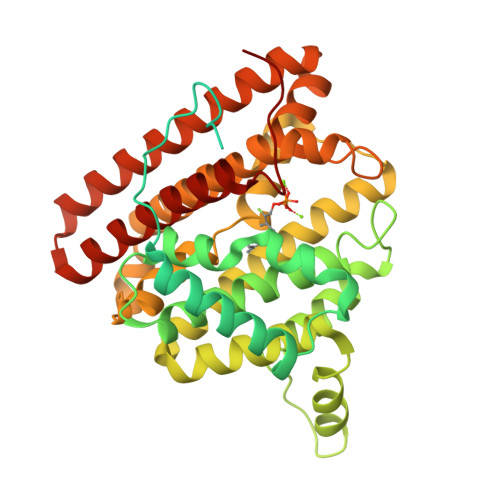
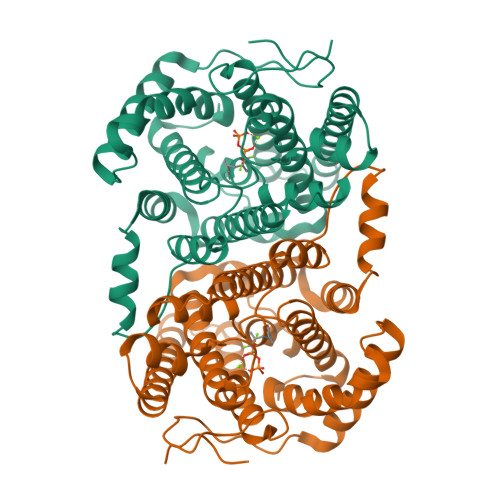
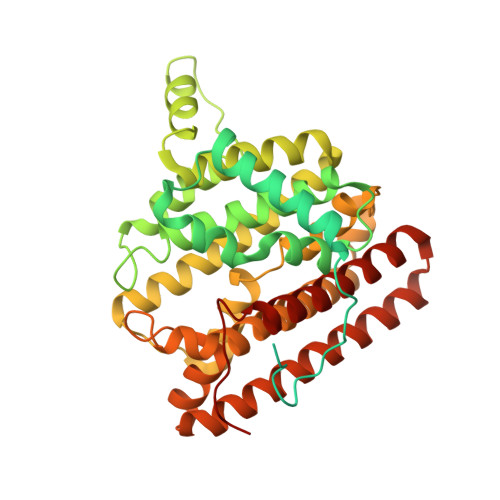
Unexpected reactivity of 2-fluorolinalyl diphosphate in the active site of crystalline 2-methylisoborneol synthase.
Koksal, M., Chou, W.K., Cane, D.E., Christianson, D.W.(2013) Biochemistry 52: 5247-5255
- PubMed: 23844678
- DOI: https://doi.org/10.1021/bi400797c
- Primary Citation of Related Structures:
4LA5, 4LA6 - PubMed Abstract:
The crystal structure of 2-methylisoborneol synthase (MIBS) from Streptomyces coelicolor A3(2) has been determined in its unliganded state and in complex with two Mg(2+) ions and 2-fluoroneryl diphosphate at 1.85 and 2.00 Å resolution, respectively. Under normal circumstances, MIBS catalyzes the cyclization of the naturally occurring, noncanonical 11-carbon isoprenoid substrate, 2-methylgeranyl diphosphate, which first undergoes an ionization-isomerization-ionization sequence through the tertiary diphosphate intermediate 2-methyllinalyl diphosphate to enable subsequent cyclization chemistry. MIBS does not exhibit catalytic activity with 2-fluorogeranyl diphosphate, and we recently reported the crystal structure of MIBS complexed with this unreactive substrate analogue [ Köksal, M., Chou, W. K. W., Cane, D. E., Christianson, D. W. (2012) Biochemistry 51 , 3011-3020 ]. However, cocrystallization of MIBS with the fluorinated analogue of the tertiary allylic diphosphate intermediate, 2-fluorolinalyl diphosphate, reveals unexpected reactivity for the intermediate analogue and yields the crystal structure of the complex with the primary allylic diphosphate, 2-fluoroneryl diphosphate. Comparison with the structure of the unliganded enzyme reveals that the crystalline enzyme active site remains partially open, presumably due to the binding of only two Mg(2+) ions. Assays in solution indicate that MIBS catalyzes the generation of (1R)-(+)-camphor from the substrate 2-fluorolinalyl diphosphate, suggesting that both 2-fluorolinalyl diphosphate and 2-methyllinalyl diphosphate follow the identical cyclization mechanism leading to 2-substituted isoborneol products; however, the initially generated 2-fluoroisoborneol cyclization product is unstable and undergoes elimination of hydrogen fluoride to yield (1R)-(+)-camphor.
Organizational Affiliation:
Roy and Diana Vagelos Laboratories, Department of Chemistry, University of Pennsylvania , 231 South 34th Street, Philadelphia, Pennsylvania 19104-6323, United States.