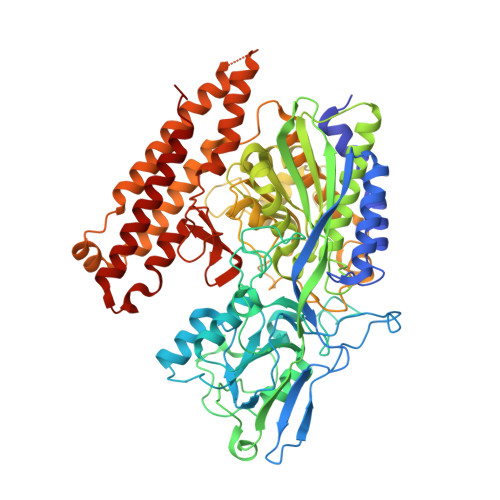
Structure-Activity Relationship of (18)F-Labeled Phosphoramidate Peptidomimetic Prostate-Specific Membrane Antigen (PSMA)-Targeted Inhibitor Analogues for PET Imaging of Prostate Cancer.
Dannoon, S., Ganguly, T., Cahaya, H., Geruntho, J.J., Galliher, M.S., Beyer, S.K., Choy, C.J., Hopkins, M.R., Regan, M., Blecha, J.E., Skultetyova, L., Drake, C.R., Jivan, S., Barinka, C., Jones, E.F., Berkman, C.E., VanBrocklin, H.F.(2016) J Med Chem 59: 5684-5694
- PubMed: 27228467
- DOI: https://doi.org/10.1021/acs.jmedchem.5b01850
- Primary Citation of Related Structures:
4LQG - PubMed Abstract:
A series of phosphoramidate-based prostate specific membrane antigen (PSMA) inhibitors of increasing lipophilicity were synthesized (4, 5, and 6), and their fluorine-18 analogs were evaluated for use as positron emission tomography (PET) imaging agents for prostate cancer. To gain insight into their modes of binding, they were also cocrystallized with the extracellular domain of PSMA. All analogs exhibited irreversible binding to PSMA with IC50 values ranging from 0.4 to 1.3 nM. In vitro assays showed binding and rapid internalization (80-95%, 2 h) of the radiolabeled ligands in PSMA(+) cells. In vivo distribution demonstrated significant uptake in CWR22Rv1 (PSMA(+)) tumor, with tumor to blood ratios of 25.6:1, 63.6:1, and 69.6:1 for [(18)F]4, [(18)F]5, and [(18)F]6, respectively, at 2 h postinjection. Installation of aminohexanoic acid (AH) linkers in the phosphoramidate scaffold improved their PSMA binding and inhibition and was critical for achieving suitable in vivo imaging properties, positioning [(18)F]5 and [(18)F]6 as favorable candidates for future prostate cancer imaging clinical trials.
Organizational Affiliation:
Department of Radiology and Biomedical Imaging, University of California-San Francisco , 185 Berry Street, San Francisco, California 94107, United States.