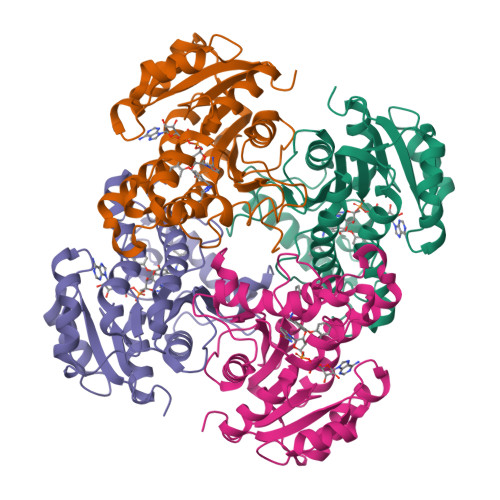
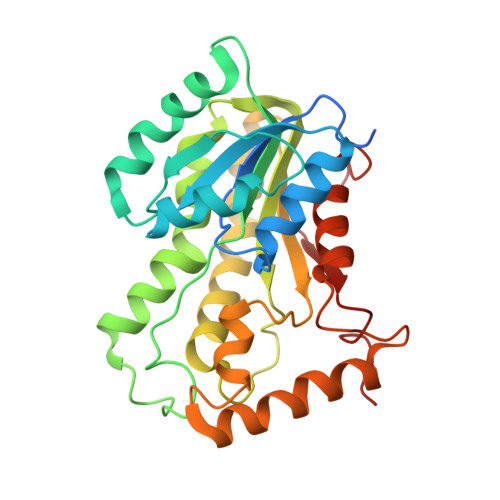
Rational Modulation of the Induced-Fit Conformational Change for Slow-Onset Inhibition in Mycobacterium tuberculosis InhA.
Lai, C.T., Li, H.J., Yu, W., Shah, S., Bommineni, G.R., Perrone, V., Garcia-Diaz, M., Tonge, P.J., Simmerling, C.(2015) Biochemistry 54: 4683-4691
- PubMed: 26147157
- DOI: https://doi.org/10.1021/acs.biochem.5b00284
- Primary Citation of Related Structures:
5COQ, 5CP8, 5CPB, 5CPF - PubMed Abstract:
Slow-onset enzyme inhibitors are the subject of considerable interest as an approach to increasing the potency of pharmaceutical compounds by extending the residence time of the inhibitor on the target (the lifetime of the drug-receptor complex). However, rational modulation of residence time presents significant challenges because it requires additional mechanistic insight, such as the nature of the transition state for postbinding isomerization. Our previous work, based on X-ray crystallography, enzyme kinetics, and molecular dynamics simulation, suggested that the slow step in inhibition of the Mycobacterium tuberculosis enoyl-ACP reductase InhA involves a change in the conformation of the substrate binding loop from an open state in the initial enzyme-inhibitor complex to a closed state in the final enzyme-inhibitor complex. Here, we use multidimensional free energy landscapes for loop isomerization to obtain a computational model for the transition state. The results suggest that slow-onset inhibitors crowd key side chains on helices that slide past each other during isomerization, resulting in a steric clash. The landscapes become significantly flatter when residues involved in the steric clash are replaced with alanine. Importantly, this lower barrier can be increased by rational inhibitor redesign to restore the steric clash. Crystallographic studies and enzyme kinetics confirm the predicted effects on loop structure and flexibility, as well as inhibitor residence time. These loss and regain of function studies validate our mechanistic hypothesis for interactions controlling substrate binding loop isomerization, providing a platform for the future design of inhibitors with longer residence times and better in vivo potency. Similar opportunities for slow-onset inhibition via the same mechanism are identified in other pathogens.