The Crystal Structure of a Class of Cyclases that Catalyze the Cope Rearrangement
Chen, C.C., Hu, X., Tang, X., Yang, Y., Ko, T.P., Gao, J., Zheng, Y., Huang, J.W., Yu, Z., Li, L., Han, S., Cai, N., Zhang, Y., Liu, W., Guo, R.T.(2018) Angew Chem Int Ed Engl 57: 15060-15064
- PubMed: 30222239
- DOI: https://doi.org/10.1002/anie.201808231
- Primary Citation of Related Structures:
5YVK, 5YVL, 5YVP, 5Z53, 5Z54, 5ZFJ, 6A8X, 6A92, 6A98, 6A99, 6A9F, 6ADU - PubMed Abstract:
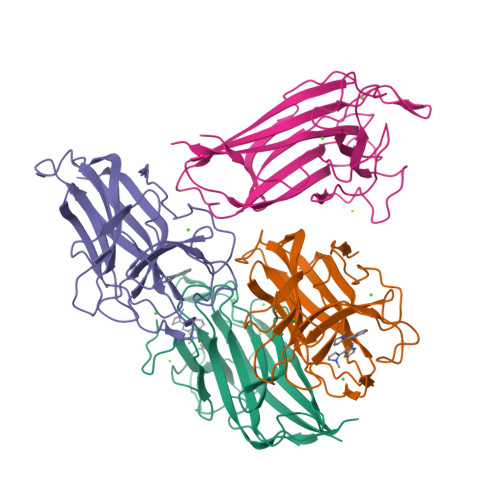
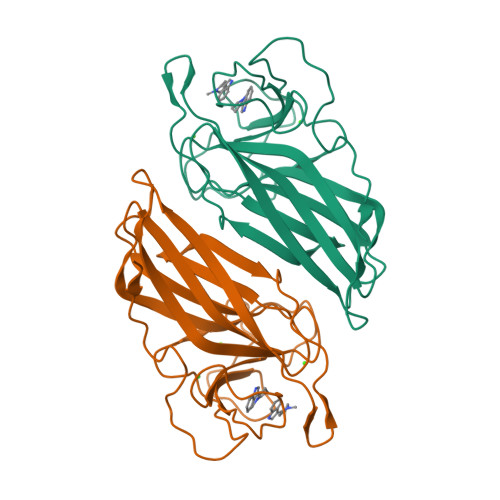
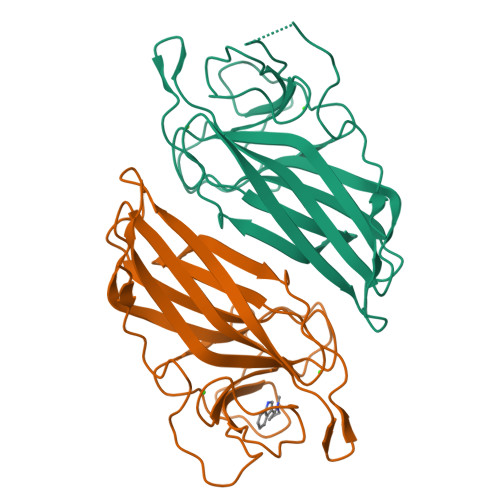
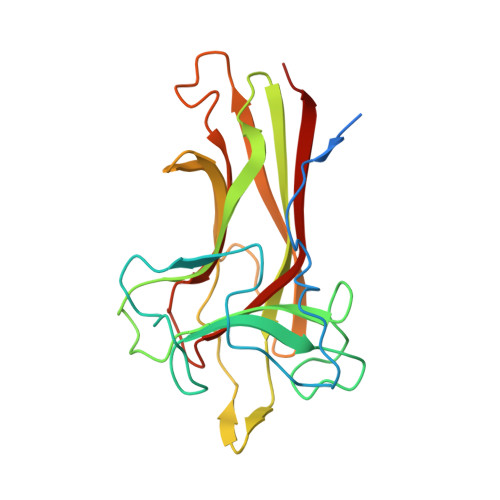
Found recently in stignomatales, the Stig cyclases catalyze the Cope rearrangement and intramolecular cyclization to produce complex indole alkaloids. Five crystal structures were solved of subfamily 1 and 2 Stig cyclases, which adopt a β-sandwich fold like the non-catalytic carbohydrate-binding motif. Several complex structures were also determined of indole-based compounds, which are bound to the hydrophobic terminal cavity, where a conserved Asp residue makes an H-bond to the indole N and triggers the acid-catalyzed Cope rearrangement. Through analyzing the enzyme-ligand interactions and mutagenesis experiments, several aromatic residues were found important in catalysis. Apart from a common substrate binding mode and catalytic mechanism, potential subfamily variations that may attribute to the different product specificity are implicated. These results shall expand our scope of enzymology, in particular for further investigation of the biosynthetic Cope rearrangement.
Organizational Affiliation:
State Key Laboratory of Biocatalysis and Enzyme Engineering, Hubei Collaborative Innovation Center for Green Transformation of Bio-resources, Environmental Microbial Technology Center of Hubei Province, Hubei Key Laboratory of Industrial Biotechnology, College of Life Sciences, Hubei University, Wuhan, 430062, China.