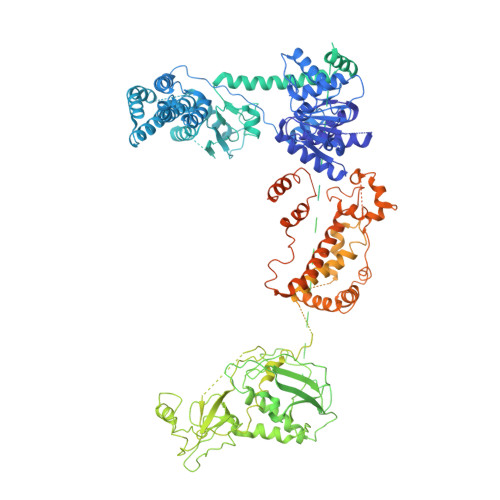
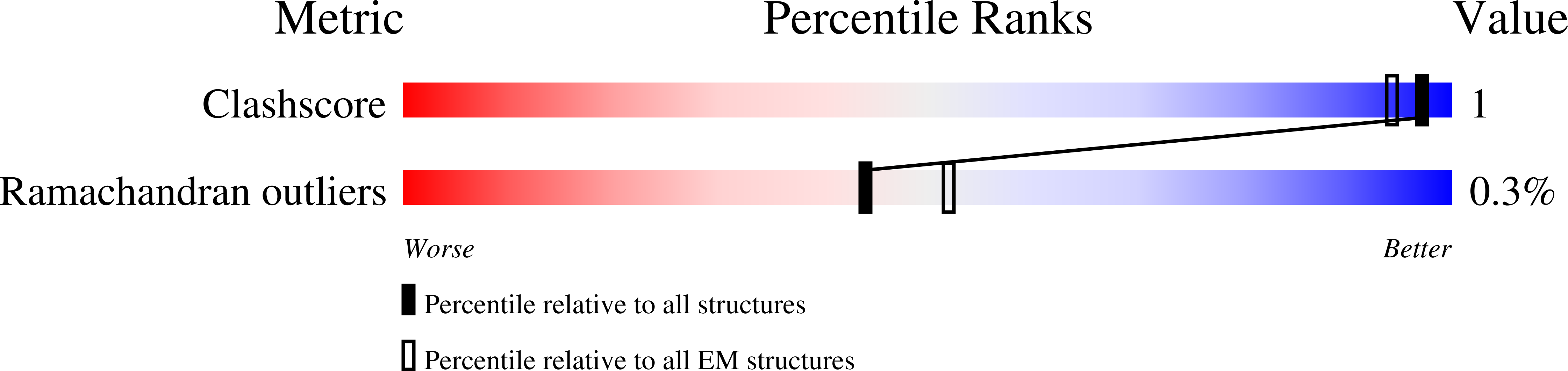Dicer uses distinct modules for recognizing dsRNA termini.
Sinha, N.K., Iwasa, J., Shen, P.S., Bass, B.L.(2018) Science 359: 329-334
- PubMed: 29269422
- DOI: https://doi.org/10.1126/science.aaq0921
- Primary Citation of Related Structures:
6BU9, 6BUA - PubMed Abstract:
Invertebrates rely on Dicer to cleave viral double-stranded RNA (dsRNA), and Drosophila Dicer-2 distinguishes dsRNA substrates by their termini. Blunt termini promote processive cleavage, while 3' overhanging termini are cleaved distributively. To understand this discrimination, we used cryo-electron microscopy to solve structures of Drosophila Dicer-2 alone and in complex with blunt dsRNA. Whereas the Platform-PAZ domains have been considered the only Dicer domains that bind dsRNA termini, unexpectedly, we found that the helicase domain is required for binding blunt, but not 3' overhanging, termini. We further showed that blunt dsRNA is locally unwound and threaded through the helicase domain in an adenosine triphosphate-dependent manner. Our studies reveal a previously unrecognized mechanism for optimizing antiviral defense and set the stage for the discovery of helicase-dependent functions in other Dicers.
Organizational Affiliation:
Department of Biochemistry, University of Utah, Salt Lake City, UT 84112, USA. [email protected] [email protected].