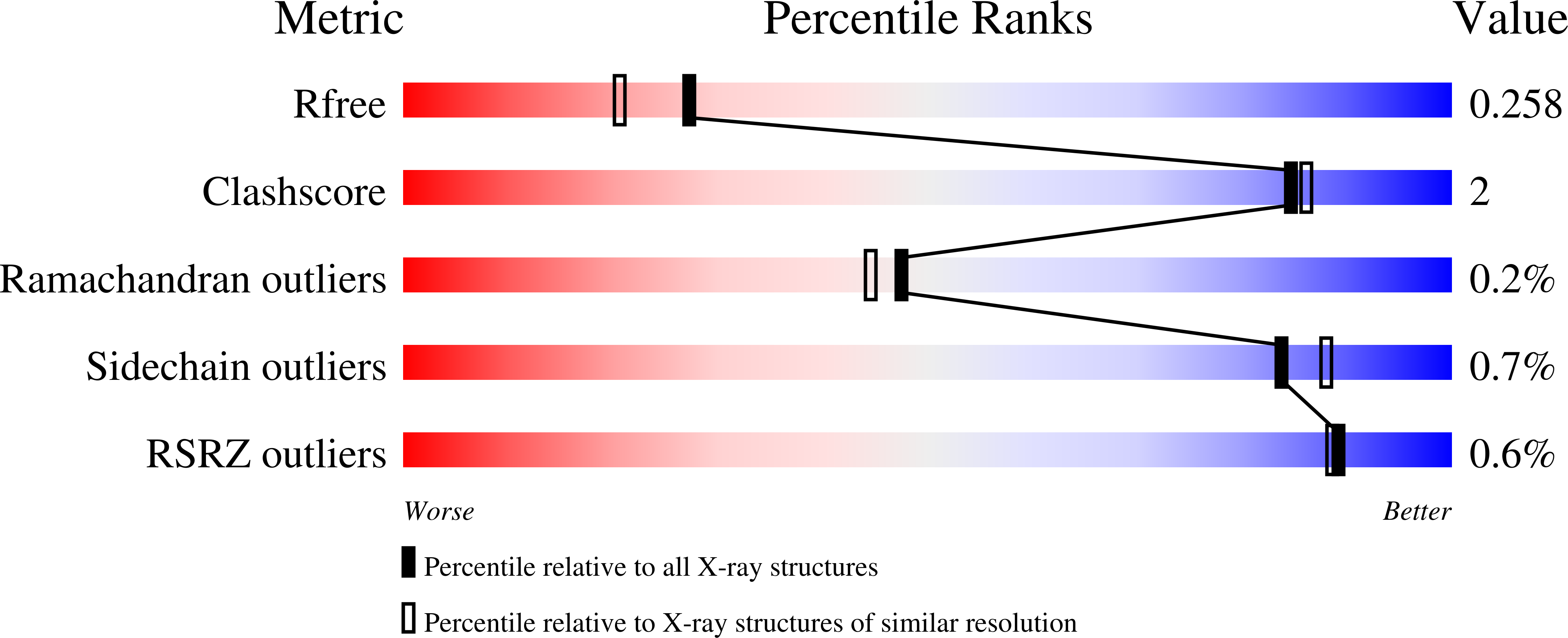
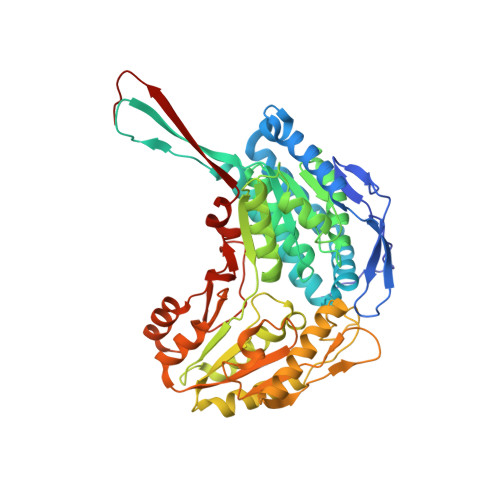
Structure-Based Optimization of a Novel Class of Aldehyde Dehydrogenase 1A (ALDH1A) Subfamily-Selective Inhibitors as Potential Adjuncts to Ovarian Cancer Chemotherapy.
Huddle, B.C., Grimley, E., Buchman, C.D., Chtcherbinine, M., Debnath, B., Mehta, P., Yang, K., Morgan, C.A., Li, S., Felton, J., Sun, D., Mehta, G., Neamati, N., Buckanovich, R.J., Hurley, T.D., Larsen, S.D.(2018) J Med Chem 61: 8754-8773
- PubMed: 30221940
- DOI: https://doi.org/10.1021/acs.jmedchem.8b00930
- Primary Citation of Related Structures:
6DUM - PubMed Abstract:
Aldehyde dehydrogenase (ALDH) activity is commonly used as a marker to identify cancer stem-like cells. The three ALDH1A isoforms have all been individually implicated in cancer stem-like cells and in chemoresistance; however, which isoform is preferentially expressed varies between cell lines. We sought to explore the structural determinants of ALDH1A isoform selectivity in a series of small-molecule inhibitors in support of research into the role of ALDH1A in cancer stem cells. An SAR campaign guided by a cocrystal structure of the HTS hit CM39 (7) with ALDH1A1 afforded first-in-class inhibitors of the ALDH1A subfamily with excellent selectivity over the homologous ALDH2 isoform. We also discovered the first reported modestly selective single isoform 1A2 and 1A3 inhibitors. Two compounds, 13g and 13h, depleted the CD133 + putative cancer stem cell pool, synergized with cisplatin, and achieved efficacious concentrations in vivo following IP administration. Compound 13h additionally synergized with cisplatin in a patient-derived ovarian cancer spheroid model.
Organizational Affiliation:
Department of Biochemistry and Molecular Biology , Indiana University School of Medicine , Indianapolis , Indiana 46202 , United States.