Nanobody-Mediated Neutralization Reveals an Achilles Heel for Norovirus.
Koromyslova, A.D., Devant, J.M., Kilic, T., Sabin, C.D., Malak, V., Hansman, G.S.(2020) J Virol 94
- PubMed: 32321816
- DOI: https://doi.org/10.1128/JVI.00660-20
- Primary Citation of Related Structures:
6XW4, 6XW5, 6XW6, 6XW7 - PubMed Abstract:
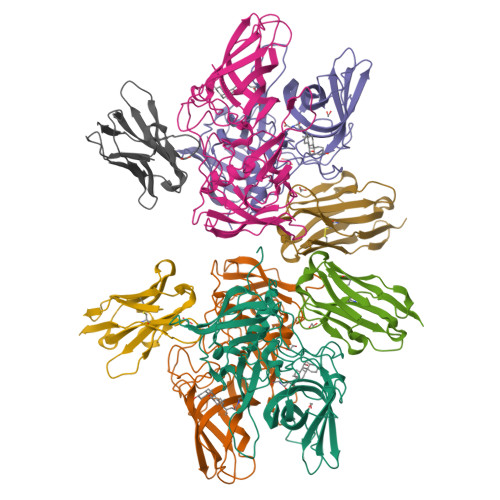
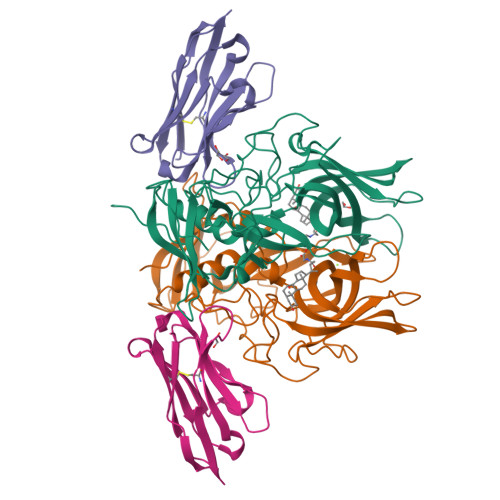
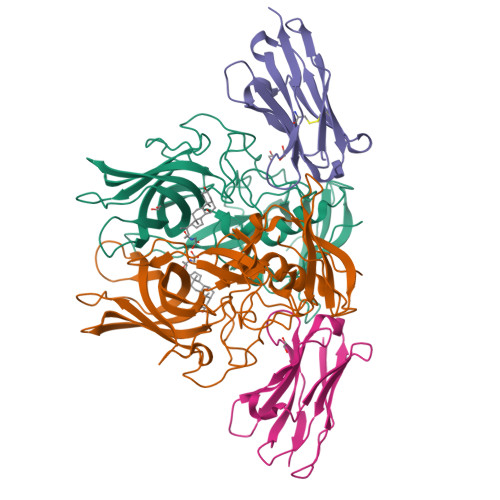
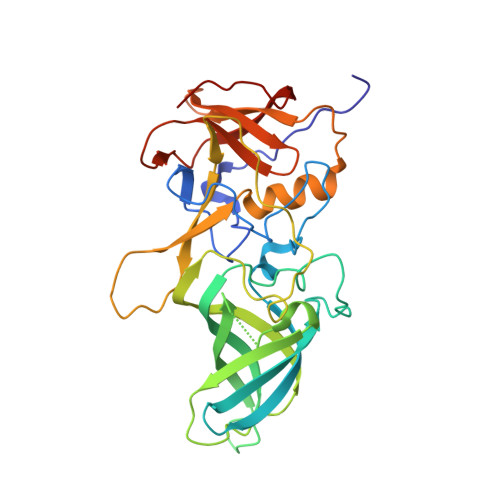
Human norovirus frequently causes outbreaks of acute gastroenteritis. Although discovered more than five decades ago, antiviral development has, until recently, been hampered by the lack of a reliable human norovirus cell culture system. Nevertheless, a lot of pathogenesis studies were accomplished using murine norovirus (MNV), which can be grown routinely in cell culture. In this study, we analyzed a sizeable library of nanobodies that were raised against the murine norovirus virion with the main purpose of developing nanobody-based inhibitors. We discovered two types of neutralizing nanobodies and analyzed the inhibition mechanisms using X-ray crystallography, cryo-electron microscopy (cryo-EM), and cell culture techniques. The first type bound on the top region of the protruding (P) domain. Interestingly, this nanobody binding region closely overlapped the MNV receptor-binding site and collectively shared numerous P domain-binding residues. In addition, we showed that these nanobodies competed with the soluble receptor, and this action blocked virion attachment to cultured cells. The second type bound at a dimeric interface on the lower side of the P dimer. We discovered that these nanobodies disrupted a structural change in the capsid associated with binding cofactors (i.e., metal cations/bile acid). Indeed, we found that capsids underwent major conformational changes following addition of Mg 2+ or Ca 2+ Ultimately, these nanobodies directly obstructed a structural modification reserved for a postreceptor attachment stage. Altogether, our new data show that nanobody-based inhibition could occur by blocking functional and structural capsid properties. IMPORTANCE This research discovered and analyzed two different types of MNV-neutralizing nanobodies. The top-binding nanobodies sterically inhibited the receptor-binding site, whereas the dimeric-binding nanobodies interfered with a structural modification associated with cofactor binding. Moreover, we found that the capsid contained a number of vulnerable regions that were essential for viral replication. In fact, the capsid appeared to be organized in a state of flux, which could be important for cofactor/receptor-binding functions. Blocking these capsid-binding events with nanobodies directly inhibited essential capsid functions. Moreover, a number of MNV-specific nanobody binding epitopes were comparable to human norovirus-specific nanobody inhibitors. Therefore, this additional structural and inhibition information could be further exploited in the development of human norovirus antivirals.
Organizational Affiliation:
Schaller Research Group at the University of Heidelberg and DKFZ, Heidelberg, Germany anna.koromyslova@gmail.com g.hansman@dkfz.de.