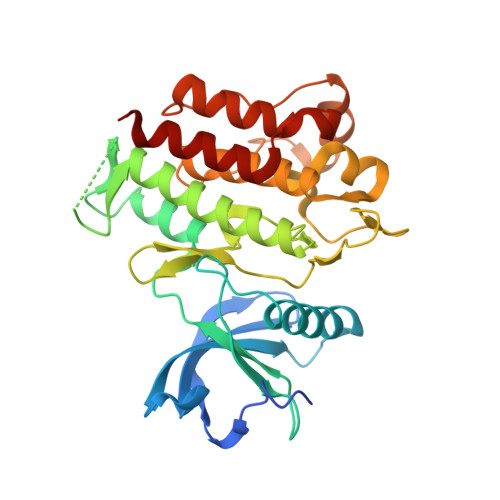
Structural basis of acquired resistance to selpercatinib and pralsetinib mediated by non-gatekeeper RET mutations.
Subbiah, V., Shen, T., Terzyan, S.S., Liu, X., Hu, X., Patel, K.P., Hu, M., Cabanillas, M., Behrang, A., Meric-Bernstam, F., Vo, P.T.T., Mooers, B.H.M., Wu, J.(2021) Ann Oncol 32: 261-268
- PubMed: 33161056
- DOI: https://doi.org/10.1016/j.annonc.2020.10.599
- Primary Citation of Related Structures:
7JU5, 7JU6 - PubMed Abstract:
Selpercatinib (LOXO-292) and pralsetinib (BLU-667) are highly potent RET-selective protein tyrosine kinase inhibitors (TKIs) for treating advanced RET-altered thyroid cancers and non-small-cell lung cancer (NSCLC). It is critical to analyze RET mutants resistant to these drugs and unravel the molecular basis to improve patient outcomes. Cell-free DNAs (cfDNAs) were analyzed in a RET-mutant medullary thyroid cancer (MTC) patient and a CCDC6-RET fusion NSCLC patient who had dramatic response to selpercatinib and later developed resistance. Selpercatinib-resistant RET mutants were identified and cross-profiled with pralsetinib in cell cultures. Crystal structures of RET-selpercatinib and RET-pralsetinib complexes were determined based on high-resolution diffraction data collected with synchrotron radiation. RET G810C/S mutations at the solvent front and RET Y806C/N mutation at the hinge region were found in cfDNAs of an MTC patient with RET M918T/V804M/L , who initially responded to selpercatinib and developed resistance. RET G810C mutant was detected in cfDNAs of a CCDC6-RET-fusion NSCLC patient who developed acquired resistance to selpercatinib. Five RET kinase domain mutations at three non-gatekeeper residues were identified from 39 selpercatinib-resistant cell lines. All five selpercatinib-resistant RET mutants were cross-resistant to pralsetinib. X-ray crystal structures of the RET-selpercatinib and RET-pralsetinib complexes reveal that, unlike other TKIs, these two RET TKIs anchor one end in the front cleft and wrap around the gate wall to access the back cleft. RET mutations at the solvent front and the hinge are resistant to both drugs. Selpercatinib and pralsetinib use an unconventional mode to bind RET that avoids the interference from gatekeeper mutations but is vulnerable to non-gatekeeper mutations.
Organizational Affiliation:
Department of Investigational Cancer Therapeutics, Division of Cancer Medicine, the University of Texas MD Anderson Cancer Center, Houston, USA. Electronic address: [email protected].