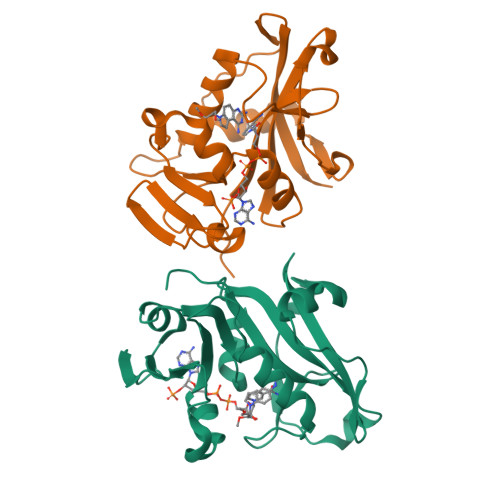
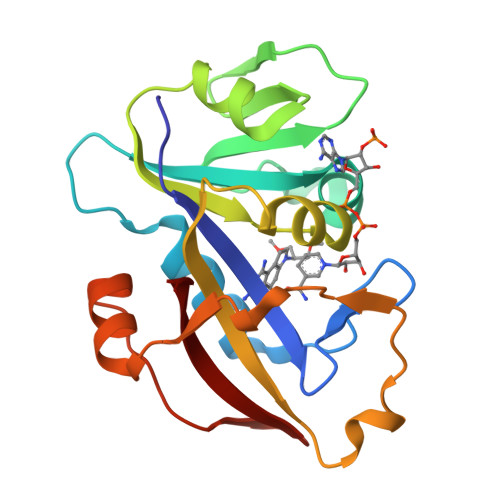
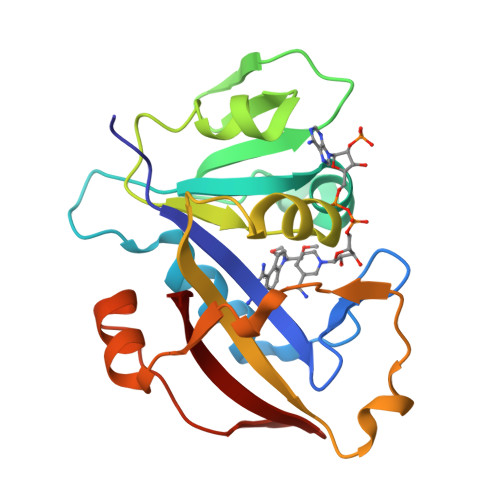
X-Ray Crystallographic Studies of Candida Albicans Dihydrofolate Reductase. High Resolution Structures of the Holoenzyme and an Inhibited Ternary Complex.
Whitlow, M., Howard, A.J., Stewart, D., Hardman, K.D., Kuyper, L.F., Baccanari, D.P., Fling, M.E., Tansik, R.L.(1997) J Biological Chem 272: 30289-30298
- PubMed: 9374515
- DOI: https://doi.org/10.1074/jbc.272.48.30289
- Primary Citation of Related Structures:
1AI9, 1AOE, 1M78, 1M79, 1M7A - PubMed Abstract:
The recent rise in systemic fungal infections has created a need for the development of new antifungal agents. As part of an effort to provide therapeutically effective inhibitors of fungal dihydrofolate reductase (DHFR), we have cloned, expressed, purified, crystallized, and determined the three-dimensional structure of Candida albicans DHFR. The 192-residue enzyme, which was expressed in Escherichia coli and purified by methotrexate affinity and cation exchange chromatography, was 27% identical to human DHFR. Crystals of C. albicans DHFR were grown as the holoenzyme complex and as a ternary complex containing a pyrroloquinazoline inhibitor. Both complexes crystallized with two molecules in the asymmetric unit in space group P21. The final structures had R-factors of 0.199 at 1.85-A resolution and 0.155 at 1.60-A resolution, respectively. The enzyme fold was similar to that of bacterial and vertebrate DHFR, and the binding of a nonselective diaminopyrroloquinazoline inhibitor and the interactions of NADPH with protein were typical of ligand binding to other DHFRs. However, the width of the active site cleft of C. albicans DHFR was significantly larger than that of the human enzyme, providing a basis for the design of potentially selective inhibitors.
Organizational Affiliation:
Genex Corporation, Protein Engineering Department, Gaithersburg, Maryland 20877, USA.