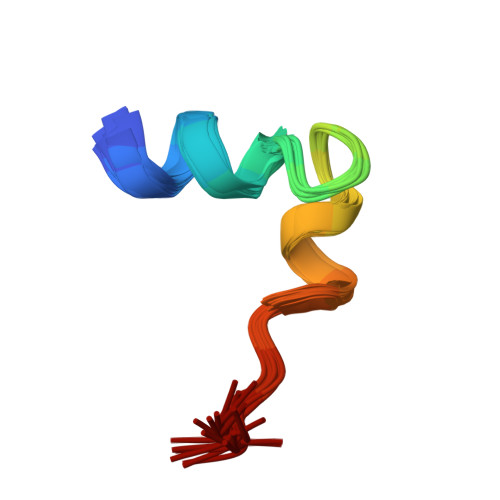
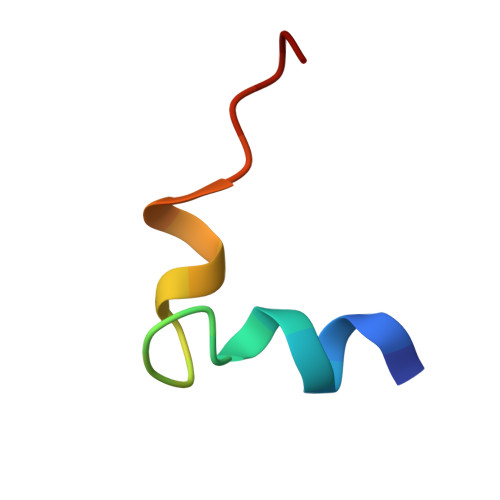
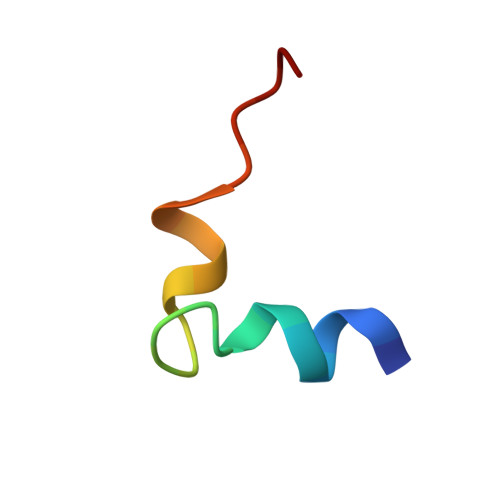
Solution conformation of an immunodominant epitope in the hepatitis B virus preS2 surface antigen.
Chi, S.W., Kim, D.H., Kim, J.S., Lee, M.K., Han, K.H.(2006) Antiviral Res 72: 207-215
- PubMed: 16872688
- DOI: https://doi.org/10.1016/j.antiviral.2006.06.009
- Primary Citation of Related Structures:
1WZ4 - PubMed Abstract:
We have determined the solution conformation of the major B cell epitope (residues 123-145, adrl23 hereafter) in the preS2 region of hepatitis B virus known to be associated with infection neutralization. The adrl23 shows an "L" shaped helix-turn-helix topology with two beta-turns formed by residues Ala(130)-Asp(133) and Asp(133)-Val(136) intervening the N- and C-terminal helices. The N-terminal alpha-helix consists of residues Ser(124)-Gln(129) whereas the C-terminal 3(10) helix is formed by residues Val(136)-Tyr(140). The beta-turns overlap partially with the putative "conformational" epitope. The overall topology of adrl23 is primarily maintained by hydrophobic interactions involving Phe(127), Leu(131), Leu(132), Val(136), and Tyr(140) that are clustered on one side of the molecule. An additional hydrophobic stabilization comes from Phe(141) that is buried inside the concave side of the molecule. A network of hydrogen bonds formed among Thr(125), His(128), and Arg(137) further contribute to the "boomerang-shaped" architecture of adrl23. The N-terminus of adrl23 is immobile due to a hydrogen bond between the N-terminal amide proton of Asn(123) and the hydroxyl oxygen of Thr(126). The side chains of Asp(133), Arg(135), Val(136), Leu(139), and Tyr(140) that were shown to be important for binding to a monoclonal antibody H8 mAb are surface exposed.
Organizational Affiliation:
Molecular Cancer Research Center, Division of Molecular Therapeutics, Korea Research Institute of Bioscience and Biotechnology, Yusong P.O. Box 115, Daejon, Korea.