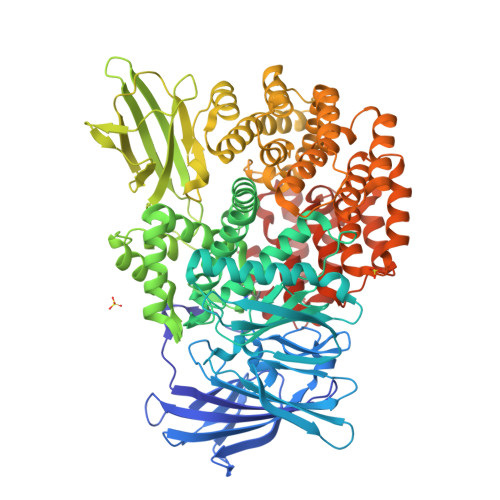
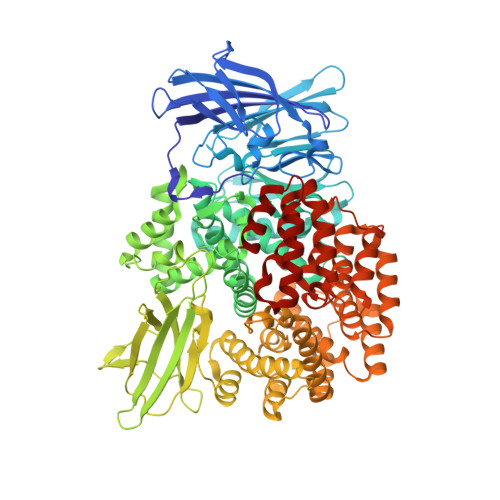
Aminopeptidase N (proteobacteria alanyl aminopeptidase) from Escherichia coli: Crystal structure and conformational change of the methionine 260 residue involved in substrate recognition
Ito, K., Nakajima, Y., Onohara, Y., Takeo, M., Nakashima, K., Matsubara, F., Ito, T., Yoshimoto, T.(2006) J Biological Chem 281: 33664-33676
- PubMed: 16885166
- DOI: https://doi.org/10.1074/jbc.M605203200
- Primary Citation of Related Structures:
2DQ6, 2DQM - PubMed Abstract:
Aminopeptidase N from Escherichia coli is a broad specificity zinc exopeptidase belonging to aminopeptidase clan MA, family M1. The structures of the ligand-free form and the enzyme-bestatin complex were determined at 1.5- and 1.6-A resolution, respectively. The enzyme is composed of four domains: an N-terminal beta-domain (Met(1)-Asp(193)), a catalytic domain (Phe(194)-Gly(444)), a middle beta-domain (Thr(445)-Trp(546)), and a C-terminal alpha-domain (Ser(547)-Ala(870)). The structure of the catalytic domain exhibits similarity to thermolysin, and a metal-binding motif (HEXXHX(18)E) is found in the domain. The zinc ion is coordinated by His(297), His(301), Glu(320), and a water molecule. The groove on the catalytic domain that contains the active site is covered by the C-terminal alpha-domain, and a large cavity is formed inside the protein. However, there exists a small hole at the center of the C-terminal alpha-domain. The N terminus of bestatin is recognized by Glu(121) and Glu(264), which are located in the N-terminal and catalytic domains, respectively. Glu(298) and Tyr(381), located near the zinc ion, are considered to be involved in peptide cleavage. A difference revealed between the ligand-free form and the enzyme-bestatin complex indicated that Met(260) functions as a cushion to accept substrates with different N-terminal residue sizes, resulting in the broad substrate specificity of this enzyme.
Organizational Affiliation:
Graduate School of Biomedical Sciences, Nagasaki University, 1-14 Bunkyo-machi, Nagasaki 852-8521, Japan.