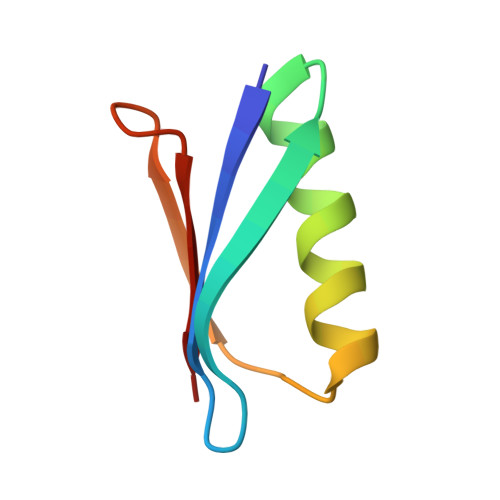
Backbone Conformational Constraints in a Microcrystalline U-15N-Labeled Protein by 3D Dipolar-Shift Solid-State NMR Spectroscopy
Franks, W.T., Wylie, B.J., Stellfox, S.A., Rienstra, C.M.(2006) J Am Chem Soc 128: 3154-3155
- PubMed: 16522090
- DOI: https://doi.org/10.1021/ja058292x
- Primary Citation of Related Structures:
2GI9 - PubMed Abstract:
Structural studies of uniformly labeled proteins by magic-angle spinning NMR spectroscopy have rapidly matured in recent years. Site-specific chemical shifts of several proteins have been assigned and structures determined from 2D or 3D data sets containing internuclear distance information. Here we demonstrate the application of a complementary technique for constraining protein backbone geometry using a site-resolved 3D dipolar-shift pulse sequence. The dipolar line shapes report on the relative orientations of 1H-15N[i] to 1H-15N[i+1] dipole vectors, constraining the torsion angles phi[i] and psi[i]. In addition, from the same 3D data set, several 1H-15N[i] to1H-15N[i+2] line shapes are extracted to constrain the torsion angles phi[i], psi[i], phi[i+1], and psi[i+1]. We report results for the majority of sites in the 56-residue beta1 immunoglobulin binding domain of protein G (GB1), using 3D experiments at 600 MHz 1H frequency. Excellent agreement between the SSNMR results and a new 1.14 A crystal structure illustrate the general potential of this technique for high-resolution structural refinement of solid proteins.
Organizational Affiliation:
Department of Chemistry, University of Illinois at Urbana-Champaign, 600 South Mathews Avenue, Urbana, Illinois 61801, USA.