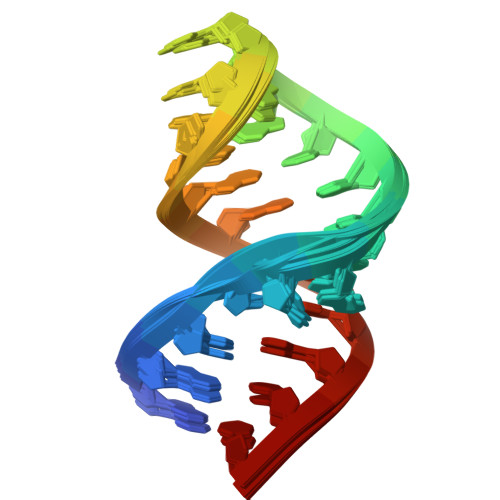
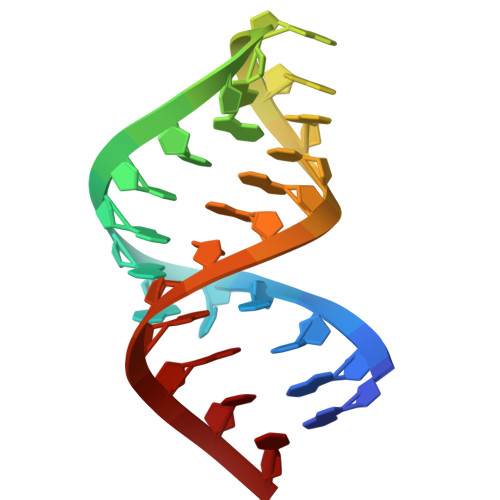
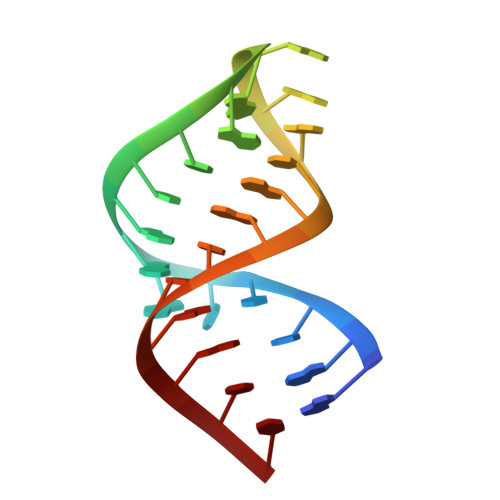
Structure of the yeast U2/U6 snRNA complex.
Burke, J.E., Sashital, D.G., Zuo, X., Wang, Y.X., Butcher, S.E.(2012) RNA 18: 673-683
- PubMed: 22328579
- DOI: https://doi.org/10.1261/rna.031138.111
- Primary Citation of Related Structures:
2LK3, 2LKR - PubMed Abstract:
The U2/U6 snRNA complex is a conserved and essential component of the active spliceosome that interacts with the pre-mRNA substrate and essential protein splicing factors to promote splicing catalysis. Here we have elucidated the solution structure of a 111-nucleotide U2/U6 complex using an approach that integrates SAXS, NMR, and molecular modeling. The U2/U6 structure contains a three-helix junction that forms an extended "Y" shape. The U6 internal stem-loop (ISL) forms a continuous stack with U2/U6 Helices Ib, Ia, and III. The coaxial stacking of Helix Ib on the U6 ISL is a configuration that is similar to the Domain V structure in group II introns. Interestingly, essential features of the complex--including the U80 metal binding site, AGC triad, and pre-mRNA recognition sites--localize to one face of the molecule. This observation suggests that the U2/U6 structure is well-suited for orienting substrate and cofactors during splicing catalysis.
Organizational Affiliation:
Department of Biochemistry, University of Wisconsin, Madison, Wisconsin 53706, USA.