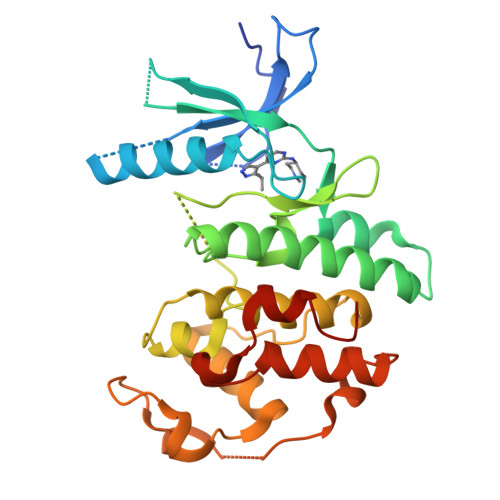
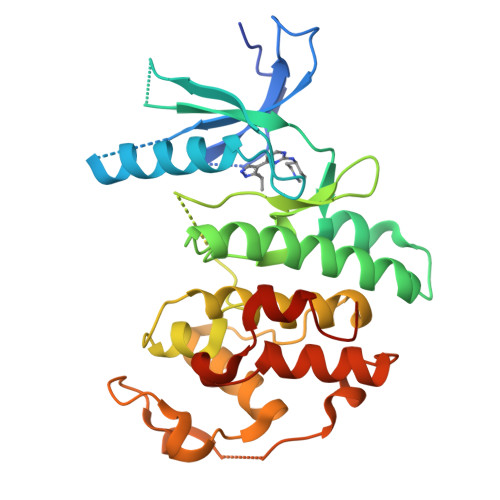
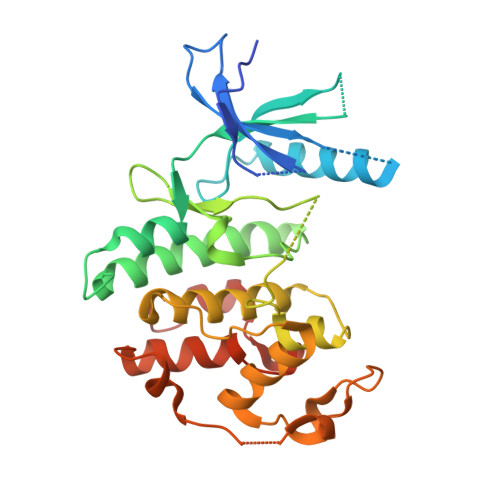
4-(Pyrazol-4-yl)-pyrimidines as selective inhibitors of cyclin-dependent kinase 4/6.
Cho, Y.S., Borland, M., Brain, C., Chen, C.H., Cheng, H., Chopra, R., Chung, K., Groarke, J., He, G., Hou, Y., Kim, S., Kovats, S., Lu, Y., O'Reilly, M., Shen, J., Smith, T., Trakshel, G., Vogtle, M., Xu, M., Xu, M., Sung, M.J.(2010) J Med Chem 53: 7938-7957
- PubMed: 21038853
- DOI: https://doi.org/10.1021/jm100571n
- Primary Citation of Related Structures:
3NUP, 3NUX - PubMed Abstract:
Identification and structure-guided optimization of a series of 4-(pyrazol-4-yl)-pyrimidines as selective CDK4/6 inhibitors is reported herein. Several potency and selectivity determinants were established based on the X-ray crystallographic analysis of representative compounds bound to monomeric CDK6. Significant selectivity for CDK4/6 over CDK1 and CDK2 was demonstrated with several compounds in both enzymatic and cellular assays.
Organizational Affiliation:
Novartis Institutes for Biomedical Research, 250 Massachusetts Avenue, Cambridge, Massachusetts 02139, USA. youngshin.cho@novartis