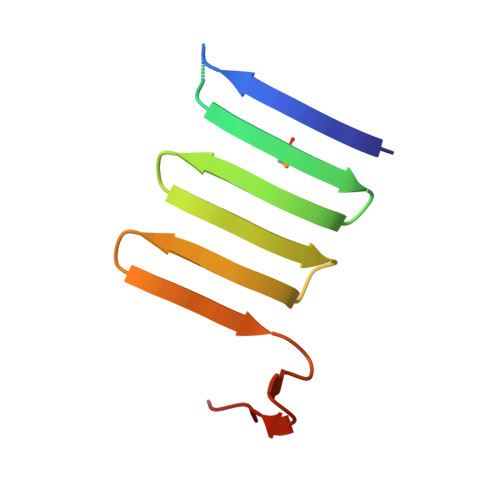
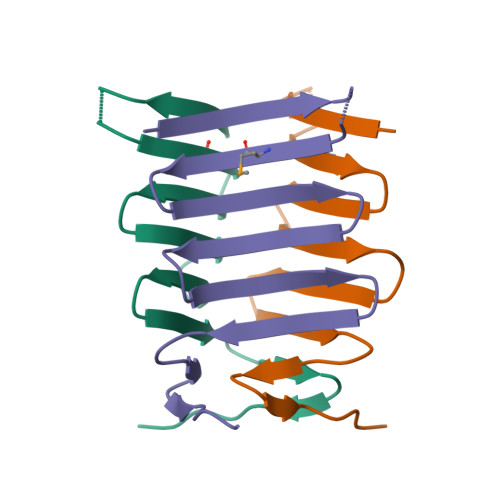
Structure and properties of the C-terminal beta-helical domain of VgrG protein from Escherichia coli O157
Uchida, K., Leiman, P.G., Arisaka, F., Kanamaru, S.(2014) J Biochem 155: 173-182
- PubMed: 24307403
- DOI: https://doi.org/10.1093/jb/mvt109
- Primary Citation of Related Structures:
3WIT - PubMed Abstract:
The bacterial Type 6 secretion system (T6SS) translocates protein toxins (also called effectors) from the cytosol of a T6SS-carrying cell to a target cell by a syringe-like supramolecular complex resembling a contractile tail of bacteriophages. Valine-glycine repeat protein G (VgrG) proteins, which are the homologues of the gp27-gp5 (gene product) cell puncturing complex of bacteriophage T4, are considered to be located at the attacking tip of the bacterial T6SS apparatus. Here, we over-expressed six VgrG proteins from pathogenic Escherichia coli O157 and CFT073 strains. Purified VgrG1 of E. coli O157 and c3393 of E. coli CFT073 form trimer in solution and are rich in β-structure. We also solved the crystal structure of a trypsin-resistant C-terminal fragment of E. coli O157 VgrG1 (VgrG1C(G561)) at 1.95 Å resolution. VgrG1C(G561) forms a three-stranded antiparallel β-helix which is structurally similar to the β-helix domain of the central spike protein (gp138) of phi92 phage, indicating a possible evolutional relationship. Comparison of four different three-stranded β-helix proteins shows how their amino acid composition determines the protein fold.
Organizational Affiliation:
Department of Bioengineering, Graduate School of Bioscience and Biotechnology, Tokyo Institute of Technology, 4259-B9 Nagatsuta-cho, Midori-ku, Yokohama 226-8501 Japan; and École Polytechnique Fédérale de Lausanne (EPFL), Laboratory of Structural Biology and Biophysics, BSP-415, CH-1015 Lausanne, Switzerland.