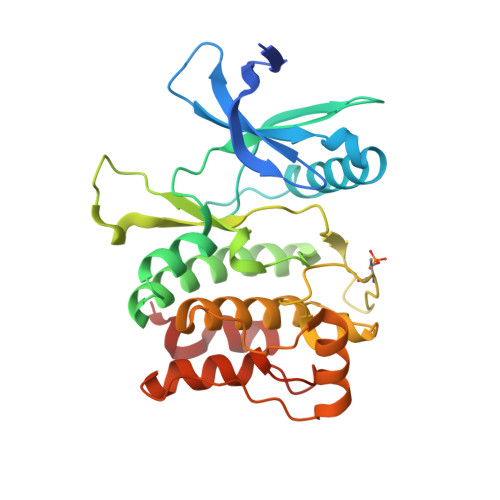
Structure of the Human Autophagy Initiating Kinase ULK1 in Complex with Potent Inhibitors.
Lazarus, M.B., Novotny, C.J., Shokat, K.M.(2015) ACS Chem Biol 10: 257-261
- PubMed: 25551253
- DOI: https://doi.org/10.1021/cb500835z
- Primary Citation of Related Structures:
4WNO, 4WNP - PubMed Abstract:
Autophagy is a conserved cellular process that involves the degradation of cellular components for energy maintenance and cytoplasmic quality control that has recently gained interest as a novel target for a variety of human diseases, including cancer. A prime candidate to determine the potential therapeutic benefit of targeting autophagy is the kinase ULK1, whose activation initiates autophagy. Here, we report the first structures of ULK1, in complex with multiple potent inhibitors. These structures show features unique to the enzyme and will provide a path for the rational design of selective compounds as cellular probes and potential therapeutics.
Organizational Affiliation:
Howard Hughes Medical Institute and Department of Cellular and Molecular Pharmacology, University of California, San Francisco , San Francisco, California 94158, United States.