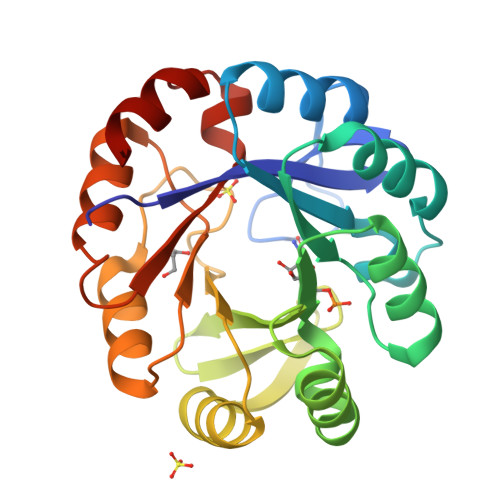
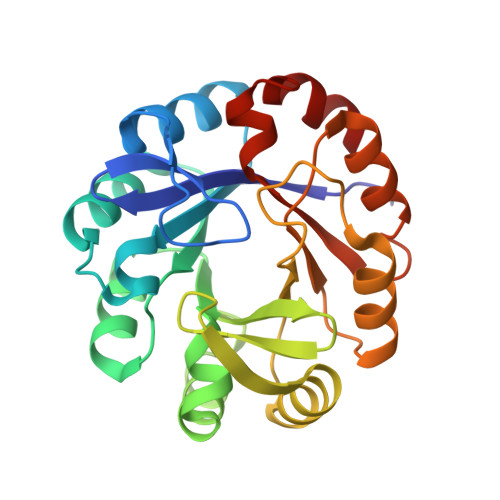
Co-occurrence of analogous enzymes determines evolution of a novel ( beta alpha )8-isomerase sub-family after non-conserved mutations in flexible loop.
Verduzco-Castro, E.A., Michalska, K., Endres, M., Juarez-Vazquez, A.L., Noda-Garcia, L., Chang, C., Henry, C.S., Babnigg, G., Joachimiak, A., Barona-Gomez, F.(2016) Biochem J 473: 1141-1152
- PubMed: 26929404
- DOI: https://doi.org/10.1042/BJ20151271
- Primary Citation of Related Structures:
4TX9, 4U28, 4W9T, 4WUI, 4X9S, 5DN1 - PubMed Abstract:
We investigate the evolution of co-occurring analogous enzymes involved in L-tryptophan and L-histidine biosynthesis in Actinobacteria Phylogenetic analysis of trpF homologues, a missing gene in certain clades of this lineage whose absence is complemented by a dual-substrate HisA homologue, termed PriA, found that they fall into three categories: (i) trpF-1, an L-tryptophan biosynthetic gene horizontally acquired by certain Corynebacterium species; (ii) trpF-2, a paralogue known to be involved in synthesizing a pyrrolopyrrole moiety and (iii) trpF-3, a variable non-conserved orthologue of trpF-1 We previously investigated the effect of trpF-1 upon the evolution of PriA substrate specificity, but nothing is known about the relationship between trpF-3 and priA After in vitro steady-state enzyme kinetics we found that trpF-3 encodes a phosphoribosyl anthranilate isomerase. However, mutation of this gene in Streptomyces sviceus did not lead to auxothrophy, as expected from the biosynthetic role of trpF-1 Biochemical characterization of a dozen co-occurring TrpF-2 or TrpF-3, with PriA homologues, explained the prototrophic phenotype, and unveiled an enzyme activity trade-off between TrpF and PriA. X-ray structural analysis suggests that the function of these PriA homologues is mediated by non-conserved mutations in the flexible L5 loop, which may be responsible for different substrate affinities. Thus, the PriA homologues that co-occur with TrpF-3 represent a novel enzyme family, termed PriB, which evolved in response to PRA isomerase activity. The characterization of co-occurring enzymes provides insights into the influence of functional redundancy on the evolution of enzyme function, which could be useful for enzyme functional annotation.
Organizational Affiliation:
Evolution of Metabolic Diversity Laboratory, Unidad de Genómica Avanzada (Langebio), Cinvestav-IPN, Irapuato, CP36821, México.