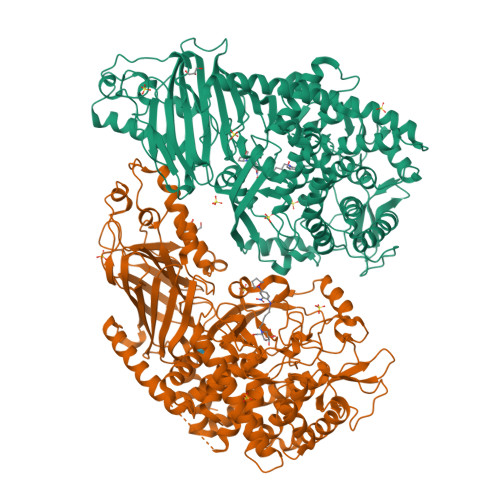
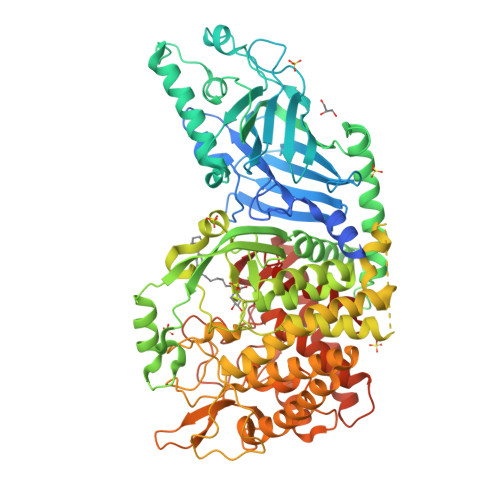
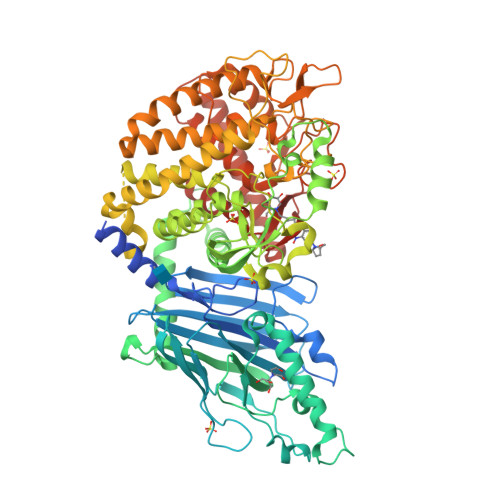
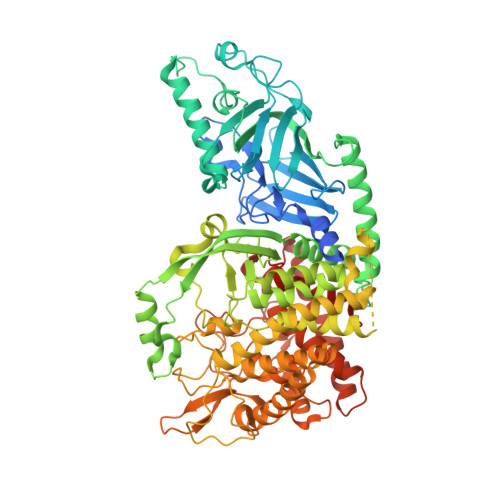
Structure-Based Design of Potent Iminosugar Inhibitors of Endoplasmic Reticulum alpha-Glucosidase I with Anti-SARS-CoV-2 Activity.
Karade, S.S., Franco, E.J., Rojas, A.C., Hanrahan, K.C., Kolesnikov, A., Yu, W., MacKerell Jr., A.D., Hill, D.C., Weber, D.J., Brown, A.N., Treston, A.M., Mariuzza, R.A.(2023) J Med Chem 66: 2744-2760
- PubMed: 36762932
- DOI: https://doi.org/10.1021/acs.jmedchem.2c01750
- Primary Citation of Related Structures:
7R6J, 7RD2, 7REV, 8E3J, 8E3P, 8E4I, 8E4K, 8E4Z, 8E5U, 8E6G, 8ECW, 8EGV, 8EHP, 8EID, 8EKN, 8ELE, 8EPJ, 8EPO, 8EPR, 8EQ7, 8EQX, 8ER4, 8ETL, 8ETO, 8EUD, 8EUR, 8EUT, 8EUX - PubMed Abstract:
Enveloped viruses depend on the host endoplasmic reticulum (ER) quality control (QC) machinery for proper glycoprotein folding. The endoplasmic reticulum quality control (ERQC) enzyme α-glucosidase I (α-GluI) is an attractive target for developing broad-spectrum antivirals. We synthesized 28 inhibitors designed to interact with all four subsites of the α-GluI active site. These inhibitors are derivatives of the iminosugars 1-deoxynojirimycin (1-DNJ) and valiolamine. Crystal structures of ER α-GluI bound to 25 1-DNJ and three valiolamine derivatives revealed the basis for inhibitory potency. We established the structure-activity relationship (SAR) and used the Site Identification by Ligand Competitive Saturation (SILCS) method to develop a model for predicting α-GluI inhibition. We screened the compounds against SARS-CoV-2 in vitro to identify those with greater antiviral activity than the benchmark α-glucosidase inhibitor UV-4. These host-targeting compounds are candidates for investigation in animal models of SARS-CoV-2 and for testing against other viruses that rely on ERQC for correct glycoprotein folding.
Organizational Affiliation:
University of Maryland Institute for Bioscience and Biotechnology Research, Rockville, Maryland 20850, United States.