Crystal structures of DCAF1-PROTAC-WDR5 complexes provide insight into DCAF1 substrate specificity
Mabanglo, M.F., Vedadi, M.To be published.
Experimental Data Snapshot
Starting Models: experimental
View more details
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| WD repeat-containing protein 5 | 329 | Homo sapiens | Mutation(s): 0 Gene Names: WDR5, BIG3 |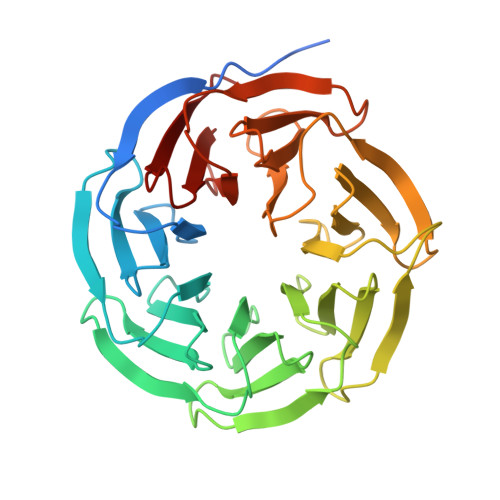  | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P61964 (Homo sapiens) Explore P61964 Go to UniProtKB: P61964 | |||||
PHAROS: P61964 GTEx: ENSG00000196363 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P61964 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| DDB1- and CUL4-associated factor 1 | 338 | Homo sapiens | Mutation(s): 0 Gene Names: DCAF1, KIAA0800, RIP, VPRBP EC: 2.7.11.1 |  | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for Q9Y4B6 (Homo sapiens) Explore Q9Y4B6 Go to UniProtKB: Q9Y4B6 | |||||
PHAROS: Q9Y4B6 GTEx: ENSG00000145041 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q9Y4B6 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| A1ANM (Subject of Investigation/LOI) Query on A1ANM | C [auth A] | N-{(1P)-5'-[(17-{(4P)-4-[(2P)-4-{[(1R)-3-amino-1-(3-chloro-4-fluorophenyl)-3-oxopropyl]carbamoyl}-3-(4-chloro-2-fluorophenyl)-1H-pyrrol-2-yl]-1H-pyrazol-1-yl}-16-oxo-3,6,9,12-tetraoxa-15-azaheptadecan-1-yl)carbamoyl]-2'-fluoro-4-[(3R,5S)-3,4,5-trimethylpiperazin-1-yl][1,1'-biphenyl]-3-yl}-6-oxo-4-(trifluoromethyl)-1,6-dihydropyridine-3-carboxamide C62 H65 Cl2 F6 N11 O10 IEXYZXCDEBBCLL-YGQKNTHVSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 79.863 | α = 90 |
| b = 83.714 | β = 90 |
| c = 130.126 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| XDS | data reduction |
| pointless | data scaling |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Ontario Institute for Cancer Research | Canada | -- |