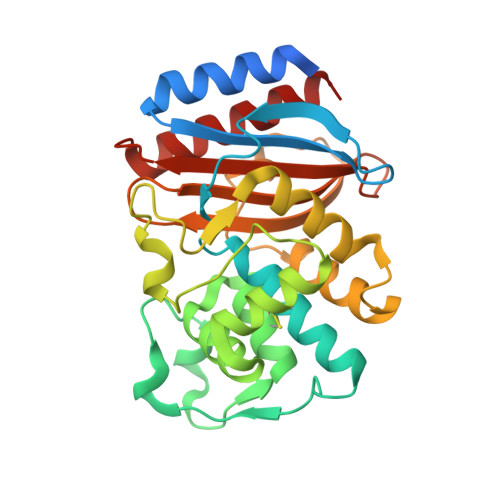
Structure of the wild-type TEM-1 beta-lactamase at 1.55 A and the mutant enzyme Ser70Ala at 2.1 A suggest the mode of noncovalent catalysis for the mutant enzyme.
Stec, B., Holtz, K.M., Wojciechowski, C.L., Kantrowitz, E.R.(2005) Acta Crystallogr D Biol Crystallogr 61: 1072-1079
- PubMed: 16041072
- DOI: https://doi.org/10.1107/S0907444905014356
- Primary Citation of Related Structures:
1ZG4, 1ZG6 - PubMed Abstract:
One of the best-studied examples of a class A beta-lactamase is Escherichia coli TEM-1 beta-lactamase. In this class of enzymes, the active-site serine residue takes on the role of a nucleophile and carries out beta-lactam hydrolysis. Here, the structures of the wild-type and the S70G enzyme determined to 1.55 and 2.1 A, respectively, are presented. In contrast to the previously reported 1.8 A structure, the active site of the wild-type enzyme (1.55 A) structure does not contain sulfate and Ser70 appears to be in the deprotonated form. The X-ray crystal structure of the S70G mutant has an altered Ser130 side-chain conformation that influences the positions of water molecules in the active site. This change allows an additional water molecule to be positioned similarly to the serine hydroxyl in the wild-type enzyme. The structure of the mutant enzyme suggests that this water molecule can assume the role of an active-site nucleophile and carry out noncovalent catalysis. The drop in activity in the mutant enzyme is comparable to the drop observed in an analogous mutation of the nucleophilic serine in alkaline phosphatase, suggesting common chemical principles in the utilization of nucleophilic serine in the active site of different enzymes.
Organizational Affiliation:
Department of Chemistry, University of Texas at El Paso, El Paso, TX 79968, USA. [email protected]