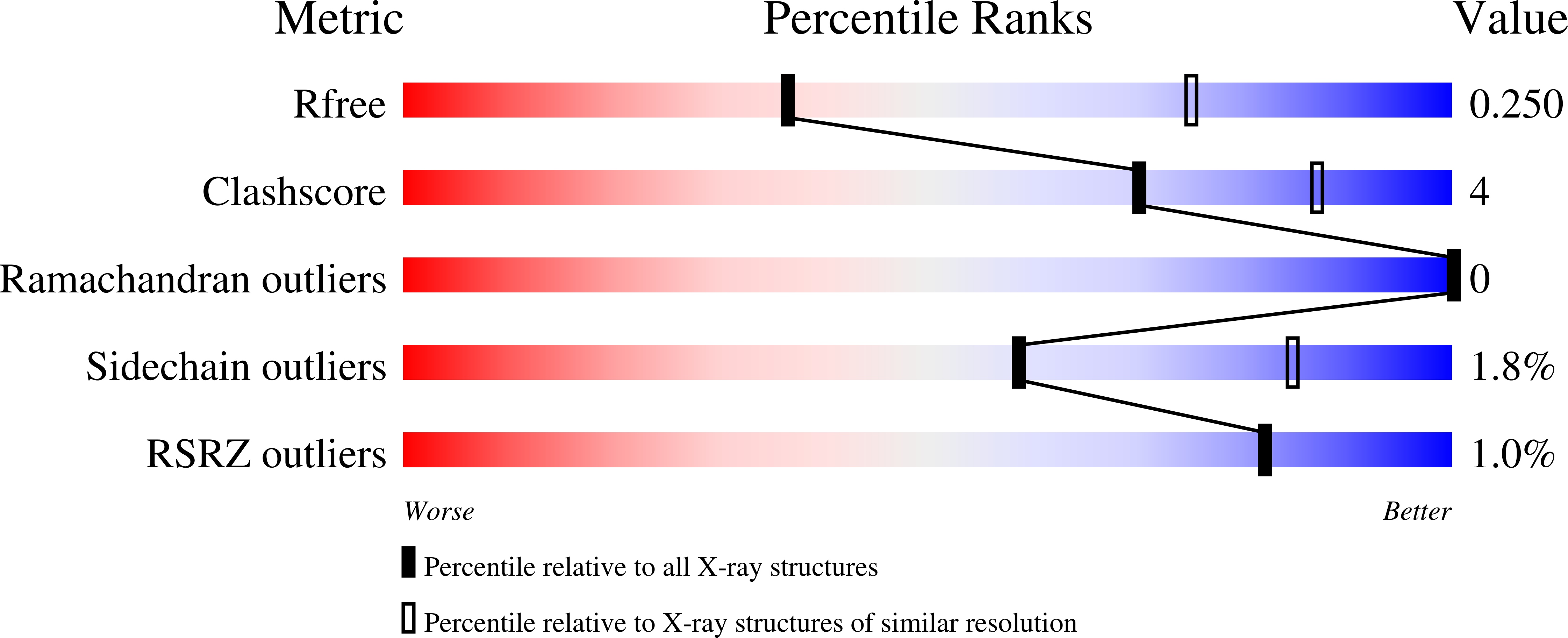
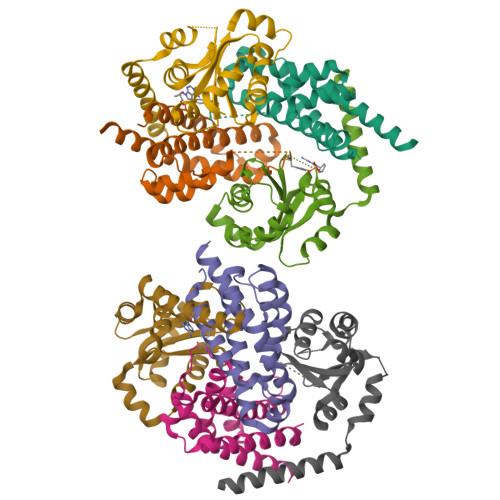
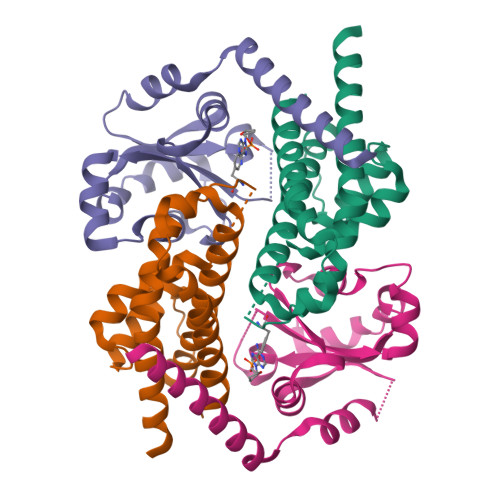
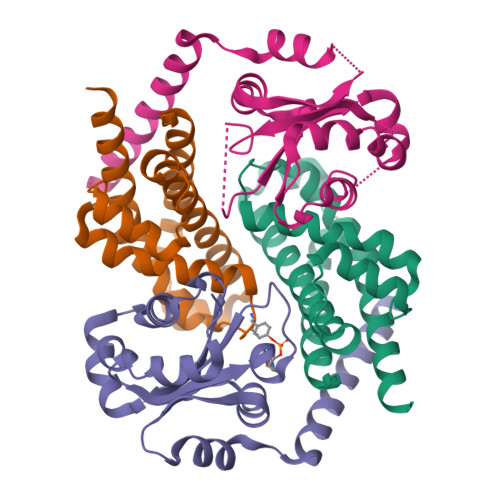
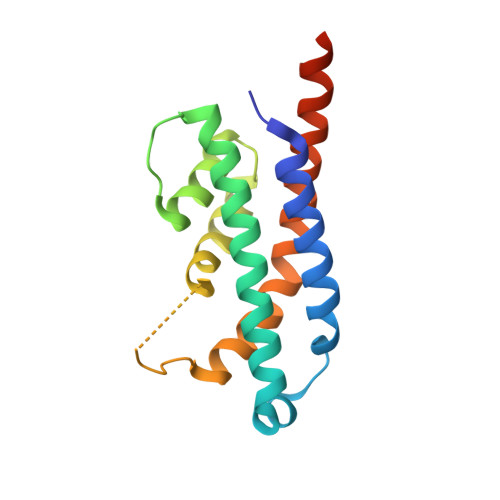
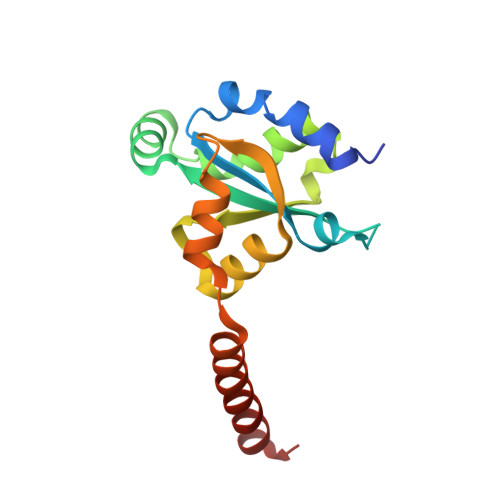
HEPN-MNT Toxin-Antitoxin System: The HEPN Ribonuclease Is Neutralized by OligoAMPylation.
Songailiene, I., Juozapaitis, J., Tamulaitiene, G., Ruksenaite, A., Sulcius, S., Sasnauskas, G., Venclovas, C., Siksnys, V.(2020) Mol Cell 80: 955-970.e7
- PubMed: 33290744
- DOI: https://doi.org/10.1016/j.molcel.2020.11.034
- Primary Citation of Related Structures:
7AE2, 7AE6, 7AE8, 7AE9, 7AER - PubMed Abstract:
Prokaryotic toxin-antitoxin (TA) systems are composed of a toxin capable of interfering with key cellular processes and its neutralizing antidote, the antitoxin. Here, we focus on the HEPN-MNT TA system encoded in the vicinity of a subtype I-D CRISPR-Cas system in the cyanobacterium Aphanizomenon flos-aquae. We show that HEPN acts as a toxic RNase, which cleaves off 4 nt from the 3' end in a subset of tRNAs, thereby interfering with translation. Surprisingly, we find that the MNT (minimal nucleotidyltransferase) antitoxin inhibits HEPN RNase through covalent di-AMPylation (diadenylylation) of a conserved tyrosine residue, Y109, in the active site loop. Furthermore, we present crystallographic snapshots of the di-AMPylation reaction at different stages that explain the mechanism of HEPN RNase inactivation. Finally, we propose that the HEPN-MNT system functions as a cellular ATP sensor that monitors ATP homeostasis and, at low ATP levels, releases active HEPN toxin.
Organizational Affiliation:
Institute of Biotechnology, Life Sciences Center, Vilnius University, Sauletekio av. 7, 10257 Vilnius, Lithuania.